Senior Investigator Research Interests
One of the fundamental rules in biology is Mendel’s Law of Segregation, which states that two alleles of a gene have an equal chance to be transmitted to gametes. Such segregation will result in the typical 3:1 phenotype ratio as in Mendel’s experiments with garden peas. However, this law can be violated by selfish genetic elements, which exploit meiosis to increase their own rate of transmission. This genetic cheating in meiosis, meiotic drive, potentially occurs whenever haploid gametes are produced from diploid parents, a process inherent to all sexually reproducing eukaryotes.
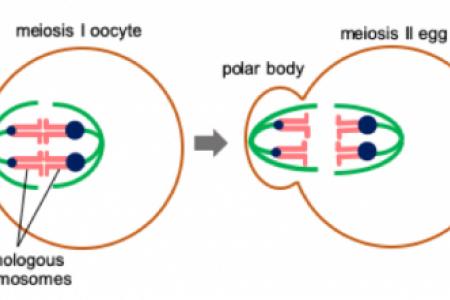
Female meiosis provides an opportunity for genetic elements to cheat because of the highly asymmetric cell division: only chromosomes segregated to the egg are transmitted to the descendants, while the rest are degraded in polar bodies. Meiotic drive impacts genetics, evolution, and fertility, as selfish elements distort transmission ratios of linked genes and manipulate gamete production, often leading to fertility issues and chromosomal disorders in offspring (e.g., Down Syndrome). Using mouse oocytes as a model, the laboratory of Chromosome Dynamics and Evolution, led by Dr. Takashi Akera focuses on the cell biological basis of meiotic drive and its consequence on mammalian reproduction.
Our work reveals how selfish DNA break Mendel’s Law, fundamentally reshaping how we understand genetic inheritance (Akera et al. Science 2017; Akera et al. Cell 2019). Most recently, we discovered that selfish DNA can actively kill eggs and embryos that do not inherit them (Clark et al. Curr Biol. 2024)—a cellular version of, “You can’t live without me.” This mechanism ensures only offspring with selfish DNA survive, causing inheritance bias and fertility issues.
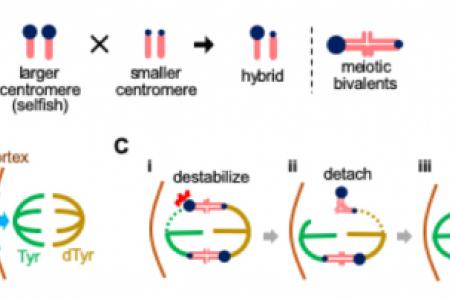
Another project explores how selfish DNA-driven genome evolution contributes to reproductive isolation. Natural variation between populations can result in hybrid sterility and lethality—hallmarks of speciation. We launched a high-risk high-reward project to test whether selfish DNA-driven genome evolution could form species barriers. We found that rapid centromere evolution causes defective chromosome dynamics in oocytes from hybrid mice, reducing female fertility and establishing reproductive barriers (El Yakoubi and Akera, Nature 2023; El Yakoubi et al., Sci Adv. 2026). Our study provides the first cell biological mechanism by which selfish DNA may drive mammalian speciation by impinging on female fertility.
While meiotic drive has been studied since 1942, our unique contribution lies in applying cutting-edge cell biology tools to visualize and manipulate selfish DNA in real time. We combine CRISPR/Cas9 system, high-resolution confocal microscopy, and optogenetic tools to capture cheating in action. Investigating meiotic drive also deepen our understanding of meiosis itself—an inherently error-prone process in women with direct relevance to fertility (Totsuka et al. Curr Biol. 2025). Roughly 15% of couples have trouble conceiving across the world, and many infertility issues may stem from genetic conflict. Our work reframes classical genetics through the lens of conflict and competition within the genome and opens up new paths to address infertility, chromosomal disorders, and speciation.
Watch a rigged game of tug-of-war inside the egg cells of mammals.
Meet the Team

Takashi Akera, Ph.D.
Dr. Takashi Akera graduated from the University of Tokyo with a Ph.D. in Biophysics and Biochemistry in 2014. He conducted postdoctoral research at the University of Pennsylvania in the laboratory of Dr. Michael Lampson from 2015 to 2019. He received both Holtzer Award for outstanding postdoctoral research in Cell and Developmental Biology and Kaushal Award for excellence in postdoctoral research in Genetics from the University of Pennsylvania in 2018. Dr. Akera joined the NHLBI in 2019 as an Earl Stadtman tenure-track Investigator and has made fundamental advances in the understanding of genetic cheating and its impact on mammalian reproduction and speciation. He has received the 2024 NHLBI Outstanding Mentor Award and the 2026 Vilcek Prize for Creative Promise in Biomedical Sciences.
Contact the lab

Warif El Yakoubi, PhD

Zaak Walton

Takaya Totsuka, Ph.D.

Duílio Silva, PhD

Sebastian Khan

Morgan Skinner

Preethi Kunchala, Ph.D.

Amelie Yanes
Alumni
Betsy Clark, PhD
Eddie (Bo) Pan, PhD
Lab Life
 Lab microinjection system |  Lab space, 2022 |
 Spring 2022; 1st lab staff photo without masks |  Fall 2023; soccer, celebrating lab's 1st paper |
 Summer 2023; lab lunch |  2026 PhD acceptance celebration lab happy hour |